Exercise¶
Introduction to BLAST using human leptin
| Authors: | Justin R. DiAngelo, Alexis Nagengast, Wilson Leung |
|---|---|
| Last Update: | Aug 31, 2019 |
| Version: | 0.0.1 |
What is BLAST?¶
BLAST stands for Basic Local Alignment Search Tool and it is a program that reports regions of similarity (at the nucleotide or protein level) between a query (your input) sequence and sequences within a database. BLAST uses a robust statistical framework that determines if the alignment between two sequences is statistically significant (i.e. has a low probability of the reported alignment being produced by chance alone). The ability to detect sequence similarity allows scientists to determine if a gene or a protein is related to other known genes or proteins in the same species or between species.
The theory of evolution is based on all organisms descending from common ancestors by speciation. At the molecular level, an ancestral DNA sequence diverges over time (through accumulation of point mutations, duplications, deletions, transpositions, recombination events, etc.) to produce diverse sequences in the genomes of living organisms. Such sequences are classified as homologs if they come from the same ancestral gene.
Mutations in genes with an important biological function have a higher probability of being harmful to the organism and are less likely to become fixed in a population. Such sequences are said to be under negative selection, which causes them to be conserved against change over time. Therefore, it is expected that two homologous copies of a functional sequence will show a higher degree of sequence conservation (observed as base-by-base similarity at the nucleotide level) than either two unrelated sequences or two sequences that are not under strong negative selection. This similarity is the “signal” detected by a BLAST search.
Obtaining sequence using NCBI¶
Learning Objectives
- Obtain a desired DNA or Protein sequence from the NCBI public database
The National Center for Biotechnology Information (NCBI) is a public database that houses molecular biology information including sequences from thousands of different species from mammals to fungi. We will explore some of the basic functionalities of the NCBI web site using leptin (LEP) — a gene that has been found to contain mutations associated with severe obesity and the development of type 2 diabetes. First, open a web browser and navigate to the NCBI web site at https://www.ncbi.nlm.nih.gov. To get information on obesity, click on the Genetics and Medicine link on the left (Figure 1), scroll about 1/3 of the way down the page and click on the Genes and Disease link. Scroll down the page and click on the “Nutritional and Metabolic Diseases” link, then click on the “Obesity” link to find a non-technical description of the hormone leptin and its role in weight control. On the right panel, you will find links to other parts of NCBI that contain more information about this disease. For example, you can access the corresponding gene record in the OMIM database (a catalog of human genes and disorders) through the “OMIM” link.
Figure 1. Click on the “Genetics & Medicine” link to access the “Genes and Disease” database.
Question 1
Based on the information on this page, how does leptin control feeding?
To obtain the sequence for the human LEP gene, go back to the NCBI
homepage at https://www.ncbi.nlm.nih.gov/. At the top of the page, use
the pull-down menu next to “Search” and select Gene. Enter LEP homo
sapiens into the text box and click Search
(Figure 2).
Note
This search menu is also available at the top of the NCBI
Bookshelf page. Hence you could scroll to the top of the page, click on
the drop-down box to select Gene and then search for LEP homo
sapiens directly. You can also access the Entrez gene record through
the “Entrez Gene” link under the “Gene sequence” section on the right
panel.

Figure 2. Search for “LEP homo sapiens” in the NCBI Gene database
Scroll down to the “Search results” section. This search produces 54 results, with the first entry being the gene record for human leptin. The remaining results may mention leptin in their detailed summary. Even before we click on the leptin entry, we can already obtain some useful information about the leptin gene from the search results page. For example, the chromosomal location and OMIM entry number for leptin are shown.
Question 2
According to these search results, on which chromosome is the leptin gene located?
Click on the first match to the human LEP gene to learn more about this gene and its sequence. The Entrez Gene record for LEP shows lots of detailed information about gene structure and function.
The first section of the Entrez Gene record is the Summary section and it contains:
- The official symbol and name approved by HUGO Gene Nomenclature Committee (HGNC)
- Other synonyms that have been used to describe this gene
- The organism the gene is from and the lineage of that organism
- Links to external databases (e.g., HGNC and Ensembl) that contain similar information
- Link to the NCBI’s Online Mendelian Inheritance in Man (MIM) that provides comprehensive information aimed for the medical and scientific research community
- RefSeq status. The NCBI Reference Sequence Database
(RefSeq) is a comprehensive, curated database of non-redundant
sequence records. The accession number for a RefSeq record begins
with two characters, follow by an underscore. This prefix denotes
the type of sequence record:
- chromosomes: NC_
- genomic regions: NG_
- mRNA: NM_
- proteins: NP_
The “Genomic regions, transcripts, and products” section has a map of the gene with links to the sequence. The sequence record can be retrieved in many different formats, including FASTA (a simple text format which contains only the sequence — easiest for BLAST searching) or GenBank (more descriptive version with the sequence listed at the end). The leptin gene is on the positive strand of the chromosome (meaning that if the chromosome was oriented from left to right, your gene would be on the strand running from 5’ to 3’). Specifically, the leptin gene can be found at 128,241,201-128,257,629 of human chromosome 7 in the current assembly (GRCh38.p13).
Question 3
How do you know that the leptin gene is on the + strand of the chromosome it’s located on?
Question 4
The picture depicts two annotated transcripts (XM_005250340.5 and NM_000230.3). What is the difference between the RefSeq record that begins with the XM_ prefix and the record that begins with the NM_ prefix? Both mRNA records have 4 green boxes, 2 light green in color (1 on each end) and 2 dark green. What do you think each of those boxes refers to?
Tip
- Examine the “COMMENT” section of the GenBank record or the RefSeq FAQ page to determine the difference between the “NM_” and “XM_” RefSeq records.
- Click on the green boxes in the diagram to get more information on the LEP gene.
Sequences are most conserved between species at the amino acid level and this is what we will use in our BLAST searches. The protein sequences for the two annotated transcripts are available under the “NCBI Reference Sequences (RefSeq)” section. Note that there are multiple options to obtain the protein sequence for the leptin precursor, including the RefSeq protein database (NP_000221.1), the Consensus CDS (CCDS) database (CCDS5800.1), the UniProtKB/Swiss-Prot database (P41159) and the UniProtKB/TrEMBL database (A4D0Y8).
To do your own BLAST search, scroll down to the “NCBI Reference
Sequences (RefSeq)” section and click on the NP_000221.1 link.
Question 5
Why do you think you want the protein sequence as opposed to the nucleotide sequence?
This screen gives you information on the LEP protein sequence. Click on the FASTA link directly beneath the gene name at the top of the screen. This gives you the FASTA definition line (as indicated by >) followed by the one letter amino acid sequence. The definition line varies but it always begins with the accession number of the sequence record (e.g., NP_000221.1), followed by the gene name and the organism. Copy the sequence including the definition line for use in a BLAST search.
Performing a BLAST search¶
Learning Objectives
- Perform a BLAST search to identify similar sequences
- Interpret results from BLAST output and draw conclusion of significance
Open a new tab in your web browser, go back to
https://www.ncbi.nlm.nih.gov/ and click on the BLAST link in the
“Popular Resources” section on the right. There are many different
options for a BLAST search and the option you choose depends on the
sequence you have and the degree of conservation you are trying to
detect. Start by clicking on the Protein BLAST image under the “Web
BLAST” section and paste your sequence into the Enter Query Sequence
text box (Figure 3).
Alternatively, you could enter the accession number (i.e.,
NP_000221.1) into the text box. If you think you will be doing many
searches at once, you can enter a Job Title to distinguish one from the
other; if not, the FASTA definition line will be used.
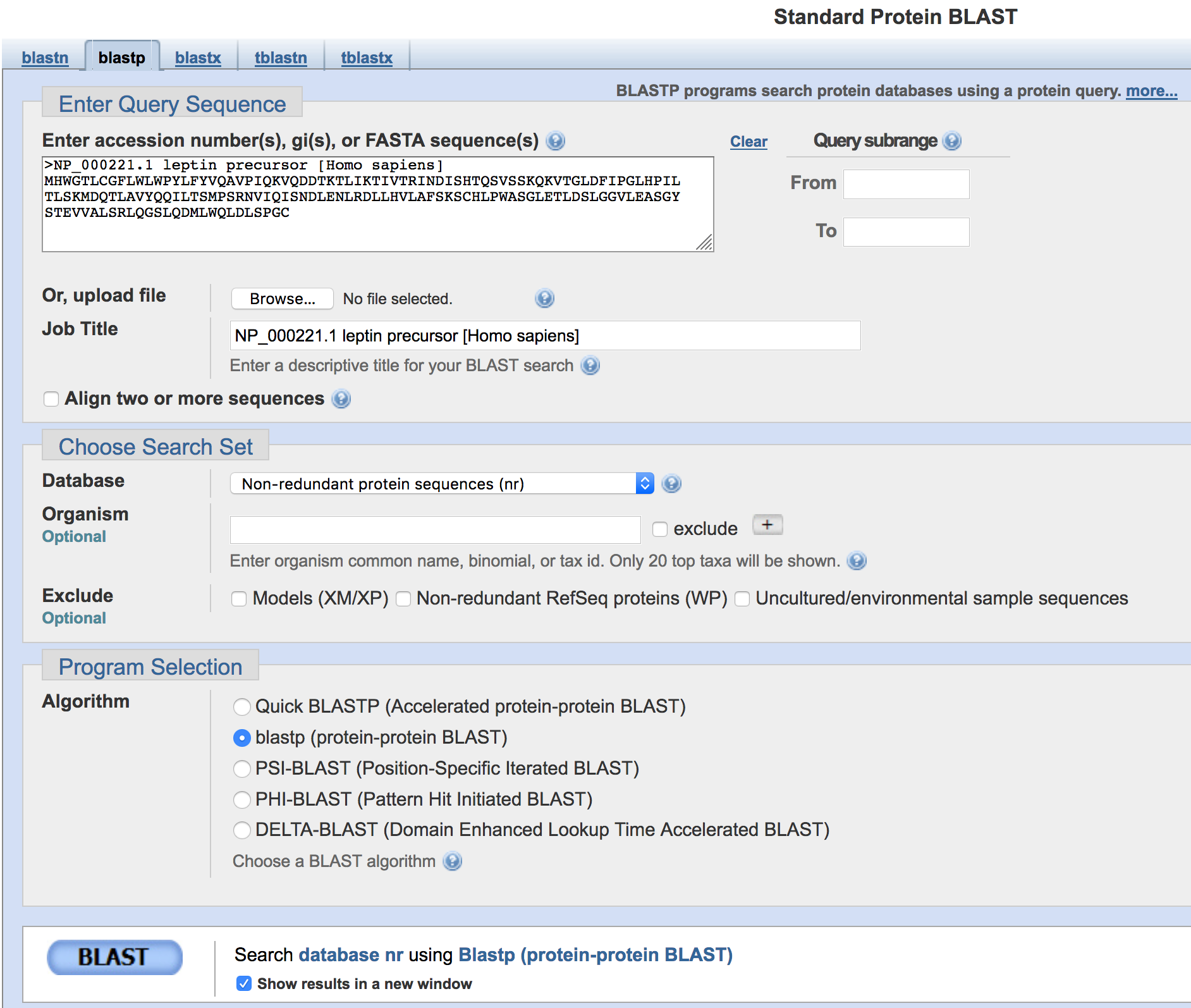
Figure 3. Configure NCBI blastp to search the human leptin protein against the nr protein database.
Archived
For teaching purposes, the result of this blastp search against the nr database has been archived and is available at the CyVerse Data Store.
Among the databases available, the non-redundant protein database is the most comprehensive; it contains sequence records from most of the other databases listed in the drop-down box:
| Database | Description |
|---|---|
| Non-redundant protein sequences (nr) | non-redundant merge of protein sequences from the NCBI’s GenBank coding sequence translations, RefSeq proteins, Swiss-Prot, PDB, PIR (Protein Information Resource at Georgetown University Medical Center), and PRF (Protein Research Foundation) |
| Reference proteins (refseq_protein) | manually curated database from NCBI |
| Model Organisms (landmark) | protein sequences from 27 genomes that cover a wide taxonomic range (from bacteria to human) |
| UniProtKB/Swiss-Prot (swissprot) | manually annotated and reviewed section of the UniProtKB database |
| Patented protein sequences (pat) | proteins from the Patent division of GenBank |
| Protein Databank proteins (pdb) | amino acid sequences derived from 3-dimensional structures from the Brookhaven Protein Data Bank |
| Metagenomic proteins (env_nr) | protein sequences from environmental samples |
| Transcriptome Shotgun Assembly proteins (tsa_nr) | transcripts assembled from ESTs and second generation sequencing reads (e.g., Illumina) |
Under the “Choose Search Set” section, select the non-redundant protein
sequences (nr) option from the “Database” drop-down menu. Then click on
the blue BLAST button to start your search. You will be taken to a page
that will update every few seconds until the BLAST search is complete.
The BLAST results page will appear once the search is complete
(Figure 4).
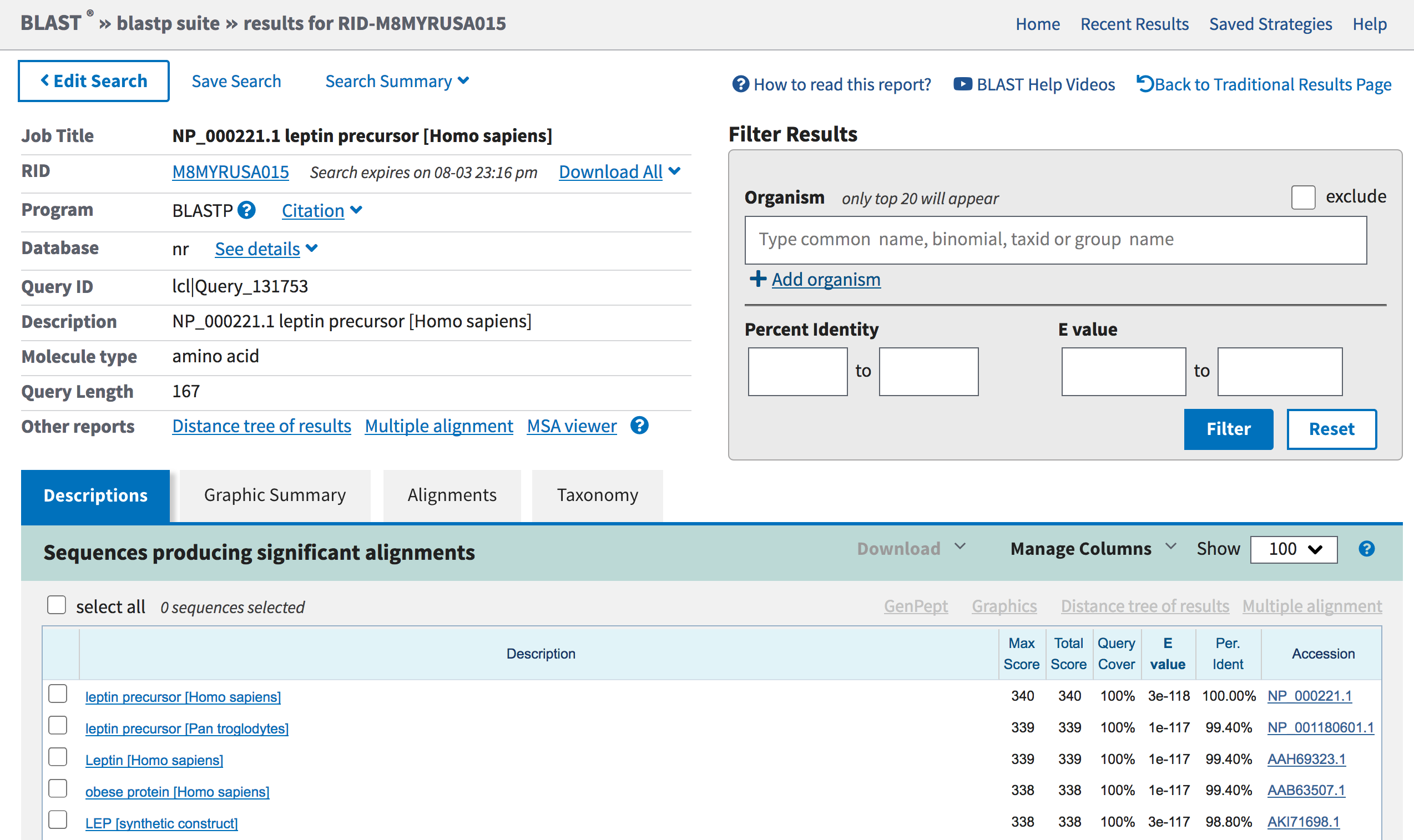
Figure 4. The search results page for the blastp search of the human leptin protein (query) against the nr database.
The top left panel provides an overview of the BLAST search parameters, and to download the BLAST results in multiple formats (e.g., Text, Hit Table). The top right panel can be used to filter the matches by organism, percent identity, and E-value. The bottom panel organizes the BLAST matches into four tabs:
- Descriptions:
- This tab contains a summary of the sequences in the database that produced significant alignments with the query sequence. Each row provides an overview of the match, including the sequence description, score, query coverage, E-value, percent identity, and accession number. Use the checkbox at the left side of each row to select or unselect a BLAST match. The other tabs will only show the information for the BLAST matches that are selected in the “Descriptions” tab.
- Graphic Summary:
- This tab shows a graphical overview of the alignments between the subject sequences in the database and the query sequence. Each row corresponds to a different subject sequence. The color of the boxes in each row correspond to the alignment score, where warmer colors (e.g., red) denote more significant matches.
- Alignments:
- This tab shows the local sequence alignments between the subject sequences in the database and the query sequence.
- Taxonomy:
- This tab provides an overview of the taxonomic distribution of the BLAST matches. Click on the buttons next to the “Reports” label to view the taxonomic groupings in three different formats. The “Lineage” report groups the matches by their taxonomic lineage (e.g., Primates). The “Organism” report groups the matches by species. The “Taxonomy” report is similar to the “Lineage” report, but you can use the “-” and “+” icons next to the non-leaf nodes in the “Taxonomy” column to collapse or expand the node.
Now that we are familiar with the different sections of the BLAST
results page, we will examine the blastp search results for the human
LEP protein sequence (query) against the nr protein database (subjects).
Click on the Graphic Summary tab to obtain a graphical overview of the
search result. Before performing the blastp search, NCBI BLAST will
search the query sequence against the Conserved Domains Database (CDD).
If the protein sequence contains a conserved domain, a graphical
representation of the alignment between the query sequence and the
conserved domain will appear at the top of the “Graphic Summary” tab.
Click on the Leptin conserved domain match in the graphic to learn more
about this conserved domain. The CDD search results page shows that the
Leptin conserved domain aligns with residues 26–167 of the query
sequence.
Go back to the Graphic Summary tab of the blastp results page. The
graphical output underneath the conserved domain image shows the
database entries (subjects) that align with your input LEP sequence
(query) on a color scale. Features with highly statistically significant
alignments will appear in red in the graphical output while features
with the lowest statistical significance will appear in black. Move your
mouse so that it is above the first red bar below the scale. A tooltip
will appear which shows the sequence title. Click on the feature to see
a more detailed description of the match (e.g., score, E-value,
accession), as well as a link to navigate directly to the corresponding
alignment (Figure 5).
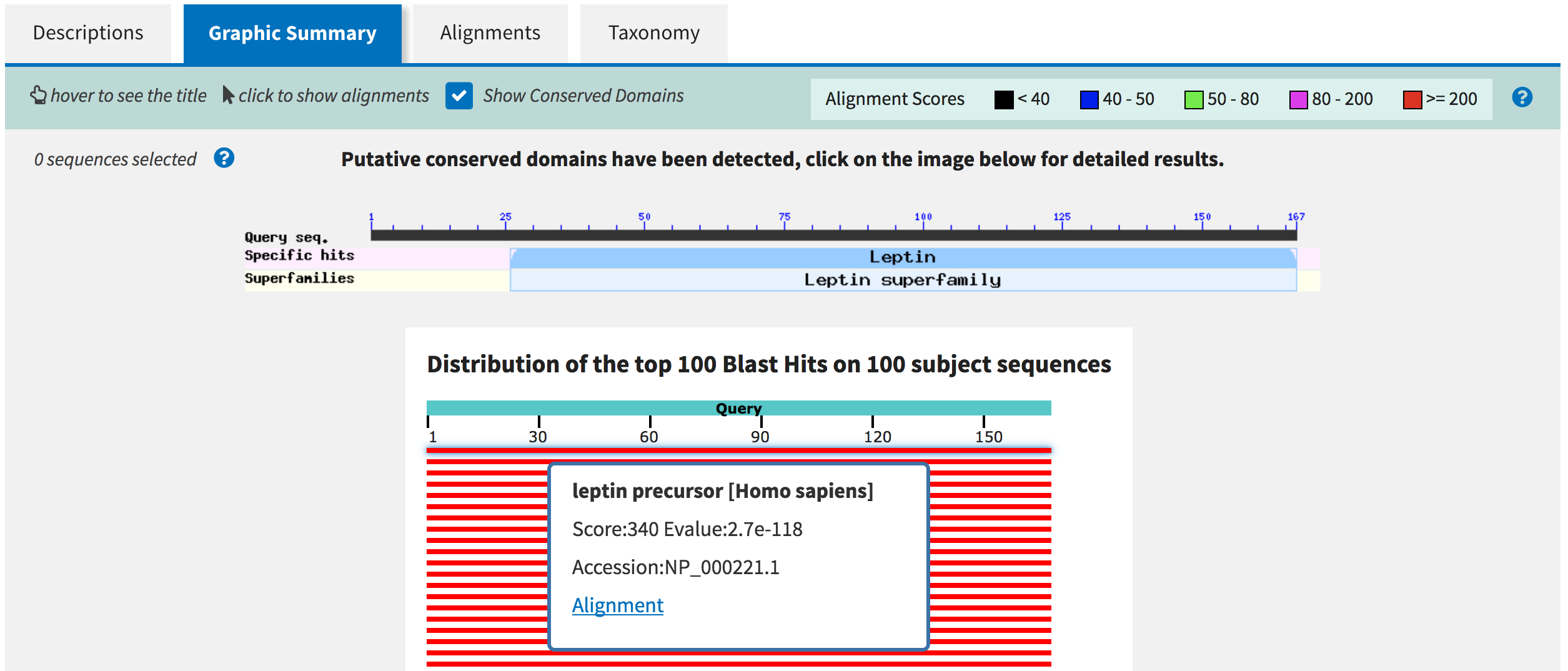
Figure 5. Click on a feature in the graphical overview diagram to obtain more information on the blastp match.
Click on the Descriptions tab to view a table summary of the blastp
matches. The first entry in the “Descriptions” table shows the best
match to the query sequence (leptin precursor [Homo sapiens]). The
match has a score (S) of 340 and an expect (E) value of 3e-118.
Alignments with smaller E-values are more statistically significant and
are less likely to have arisen by chance. Specifically, the E-value
denotes the number of times we expect to see an alignment with scores
equal to or greater than S when we align random sequences against each
other. By default, NCBI BLAST will report matches with an E-value of
less than 10. The last column of the “Descriptions” table contains the
accession number for the match (NP_000221.1), and the link allows you to
access the corresponding GenPept record.
The second entry in the “Descriptions” table shows a match to the leptin precursor in chimpanzee (Pan troglodyte). This match has a total score of 339 and an E-value of 1e-117. Because the accession number for this match (NP_001180601.1) begins with “NP_”, this protein record is from the RefSeq database.
Click on the leptin precursor [Pan troglodytes] link under the
“Description” column to navigate to the alignment between your input
leptin sequence (query) and the leptin precursor in chimp (subject). The
header in the alignment output shows that the chimp leptin precursor
sequence has a total length of 167 amino acids. The alignment has an
E-value of 1e-117 and there is a single amino acid substitution between
the human and chimp leptin precursors (from V to M). This high degree of
sequence similarity is unlikely to be caused by chance alone.
The alignment is shown as three separate lines; the first (Query) line is our human sequence, the third (Sbjct) line is the chimpanzee sequence and the middle (Match) line represents a comparison of the two sequences (if the amino acids are the same between the two sequences, that amino acid will be included in the middle line). The value in the “Identities” field corresponds to the number of identical amino acids between the query and the subject sequences. In this case, this field (166/167) indicates that, of the 167 amino acids in the alignment between the human and chimp sequences, 166 of the amino acids (99%) are identical. The one difference between these two sequences can be found at position 94 of the query (as a + in the Match line), where the human sequence is a valine (V) and the chimp sequence is a methionine (M).
Question 6
What is the relationship between V and M that warranted a + designation?
Click on the Descriptions tab in the blue toolbar. To investigate
protein matches to water buffalo (with the scientific name Bubalus
bubalis), enter Bubalus bubalis (taxid:89462) into the “Organism”
text box under the “Filter Results” section and then click on the
Filter button.
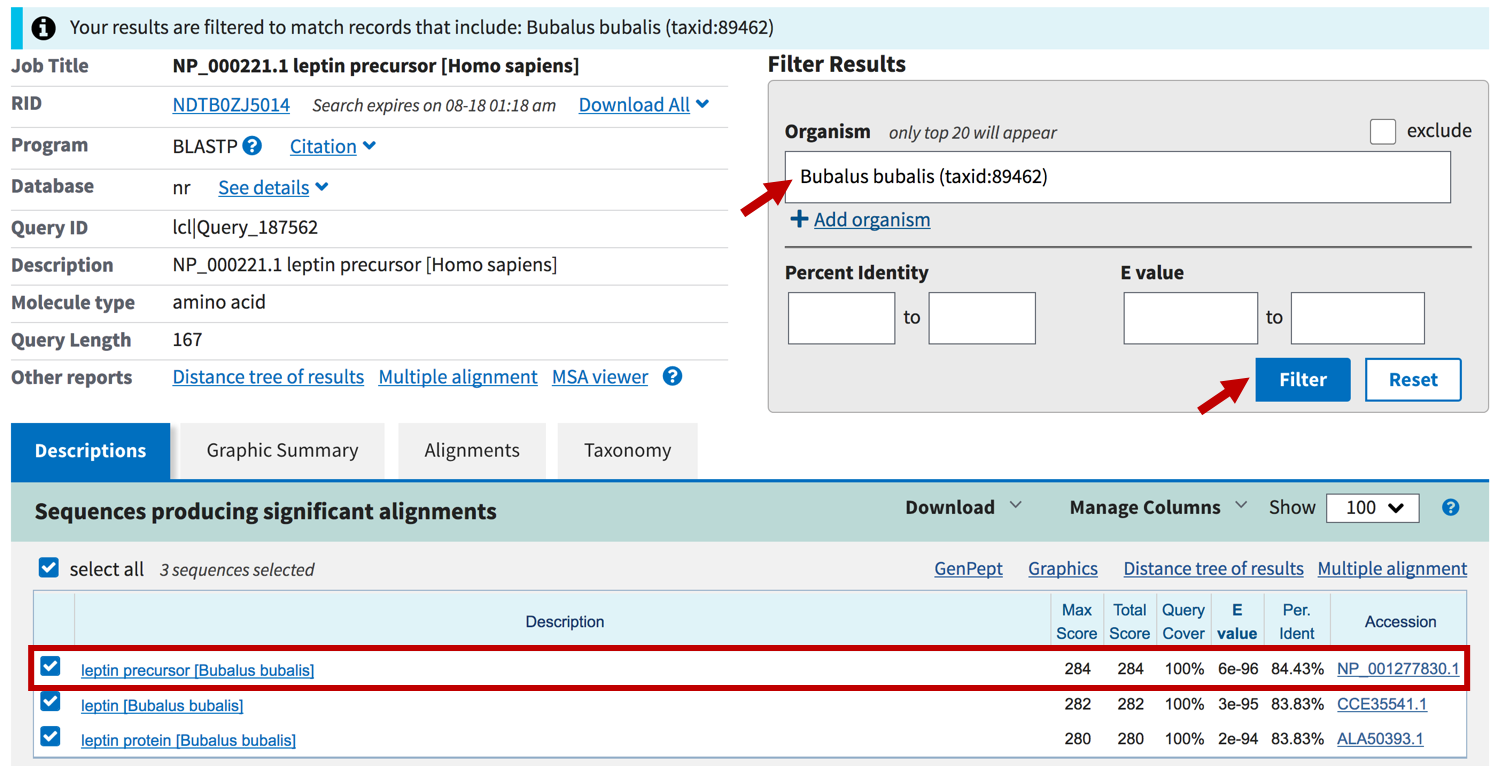
Figure 6. Identify the best match to the human leptin protein in water buffalo.
The best RefSeq match is to the leptin precursor in Bubalus bubalis with the accession number NP_001277830.1. The E-value is still very highly significant at 6e-96 (Figure 6). Click on the link under the “Description” column to navigate to the alignment. The Bubalus bubalis leptin precursor protein has the same length (167aa) as the human leptin precursor protein, and the entire protein is included in the alignment. However, there are many more changes and the percent identity are substantially lower than the alignment to the chimp leptin precursor protein.
Question 7
What is the percent identity and percent positives (aka percent similarity) of the Bubalus bubalis leptin sequence compared to the human sequence?
You can also perform searches against the database of specific
organisms. Go to the BLAST page at
https://blast.ncbi.nlm.nih.gov/Blast.cgi and enter the name of the
organism in the text box under the “BLAST Genomes” section to search its
databases. For example, enter chimpanzee (taxid:9598) into the text
box and then click on the Search button to search the chimp assembly.
To search for protein matches, click on the blastp tab, paste the
human leptin precursor sequence into the Enter Query Sequence text
box, and then verify that the RefSeq protein option is selected under
the “Database” field.
Because the database is much smaller (with 64,958 sequences) than the nr database, this search will run much faster than our previous search. Click on the BLAST button to begin your search.
Archived
For teaching purposes, the result of this blastp search against the chimpanzee RefSeq protein database has been archived and is available at the CyVerse Data Store.
Question 8
How many hits came out of this search? Are all of them significant matches? Does the BLAST result support the hypothesis that chimps have a homolog of the human leptin gene?
Tip
In general, we consider matches with E-values less than 1 x 10-5 as statistically significant.
To further support your hypothesis that chimps have a homolog of the
human leptin precursor gene, you can use the putative chimp leptin
precursor sequence and do a blastp search against the human RefSeq
protein sequence database. To do this, select the Alignments tab and
then click on the GenPept link in the blue toolbar above the sequence
you believe to be the chimp leptin precursor to retrieve the GenPept
protein record. Click on the FASTA link under the gene name to obtain
the full chimp protein sequence in FASTA format. Copy the sequence and
go back to the NCBI BLAST page and click on the Human link under the
“BLAST Genomes” section. Select the blastp program, verify that the
RefSeq protein option is selected under the “Database” field, and then
paste the chimp leptin precursor sequence into the Enter Query
Sequence text box.
Click on the BLAST button to perform your search.
Archived
For teaching purposes, the result of this blastp search against the human RefSeq protein database has been archived and is available at the CyVerse Data Store.
Question 9
Did you obtain matches from this search? Are they significant matches? How do you know? Does this data support your hypothesis about the presence of a leptin homolog in chimps?
Information accessed August 2, 2019.